Overview
The institutional review board at Tehran University of Medical Sciences passed the research protocol on 2016. Data were collected prospectively from the breast clinic at Sina Hospital, Tehran, Iran. Total study course spanned 2016-19. Written informed consent pertinent to the study type was obtained from all participants. All analysis were performed using R version 3.5.3 (R foundation for statistical computing, Vienna, Austria). Figure 1 summarizes the research sequence.
Participants
Breast clinic at Sina Hospital cares for patients with any breast related complaint mostly from lower socioeconomic levels and receives both new patients and referrals. All patients are primarily visited by junior general surgery residents who also record relevant patient information. They are then presented to senior residents or attending surgeons who shared with patients, decide the proper course of intervention. Study population included all consenting women visiting the clinic for the first time. The following were excluded: 1. Patients who had a history of breast cancer in the past; 2. Patients who had a pathological evaluation or a definite diagnosis for a breast disease from another center; 3. Patients who had a known genetic predisposition to breast cancer (such as BRCA1/2 mutation); 4. Patients who had visited the breast clinic before the commencement of this study; 5. Patients who had missing values for more than one of the selected features. 894 visiting patients were eligible for inclusion. Disease status for all eligible patients was reexamined one year after collecting the last patient data. 94 patients were lost to follow up.
Labeling outcomes
Routinely, all patients who are suspected to have a malignant breast disease undergo breast tissue biopsy and receive a pathologic evaluation by board certified pathologists at Sina Hospital. This evaluation was used as the confirmation test for positive outcomes (breast cancer) of this study; however, subjecting all patients to biopsy without indication would be unethical. Therefore the research protocol defined the reference standard for labeling outcomes as follows: Any patient with a positive biopsy for any kind of breast malignancy at any time during the study period was considered as positive. All the patients who had not undergone biopsy (i.e. had recovered from a benign disease or at the time of follow up, were still being managed under a non-malignant diagnosis) as well as patients with non-malignant biopsy specimens were considered as negative.
See the complete patient flow diagram and outcome assignment at figure 2.
Feature selection
We prioritized our specific clinical problem, availability of data to potential end-users, parsimony and overfitting avoidance 35. From the pool of routinely collected data at the clinic, we selected 17 features (independent variables) which were among the identified predictors of breast cancer in the existing literature 2, 36. They covered patient demographics, symptoms and history, findings in the clinical examination as well as mammography and ultrasound imaging (due to its common role in breast cancer screening as well as diagnosing benign breast disease 37, 38). See table 2 in the results section for a comprehensive list of all selected features.
Inputs
‘Reason for visit’ was entered in four categories: 1. Pain, discomfort, or discharge; 2. Abnormal findings in self-examination; 3.Presenting the results of mammography/ultrasound performed elsewhere; 4. asking for a second opinion on a breast related complaint (without confirmed diagnosis); 5. Other reasons. The following were entered in a binary format (1 indicating the presence and 0 indicating the absence of the condition): Breast pain (mastalgia); cyclic type of mastalgia; history of OCP (oral contraceptive pills) use; history of smoking; family history of breast disease in first degree relatives; auxiliary lymphadenopathy; breast mass; breast tenderness; breast discharge; and clinical opinion. For clinical opinion, we asked the examining physicians to mark the patients who based on history, examination and available imaging, had a high probability for malignant breast disease. Mammography/ultrasound results were also entered in a binary scale based on their reported BI-RADS scoring 39 (one indicating BI-RADS 4 and 5, and zero indicating BI-RADS 1,2 and 3). Age, age at menarche, age at first full-term pregnancy and number of full term pregnancies were entered in a numerical format.
Data preprocessing
All gathered data underwent preprocessing to optimize model training. ‘R package “missForest’ was used to impute missing values. The package uses a multivariate iterative method based on random forest algorithms. This method is non-parametric and allows for mixed data types, non-linear data structures and complex interactions 40. Using R package “mvoutlier”, a multivariate procedure that relies on robust mahalanobis distances was implemented to identify outliers 41. To remove variation in feature scales and help models converge faster, all features where normalized using the min-max scaler function:
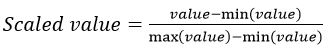
Algorithms
The purpose of this work is to train supervised algorithms for a data classification task (i.e. by training machine learning algorithms on data with known class labels, we are developing a breast cancer prediction model that classifies patients with unknown outcomes). Considering the properties of our dataset and the attributes of different machine learning and neural networks based methods 21, 42, 43, we decided to train and test six algorithms (‘associated R package’) that were most pertinent to our task: Decision tree (tree); K nearest neighbors (CORElearn); Multi-layer perceptron (RSNNS); Random Forest (randomForest); Single hidden layer (nnet); and Support vector machines (kernlab). All hyper-parameters were set as default. We implemented logistic regression to calculate the adjusted odds ratio for each independent variable.
Performance evaluation
Trained algorithms were run on all collected data. To compare performances, we calculated accuracy, sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV) and estimated prevalence with confidence intervals for each model. We provided receiver operating characteristic (ROC) curves with associated area under the curve (AUC) measurements as well. 44, 45. See the supplementary material for information on performance metrics.
Validation
Models trained on relatively small datasets are prone to bias and overfitting. Therefore in addition to the primary evaluation, we performed a 10-fold stratified cross validation for all six algorithms 46. The dataset was randomly divided into ten equal subsets. Each subset was used as a testing set, while the algorithms were trained on the remaining nine subsets. The mean and range of performance statistics for these ten iterations were reported and compared to the primary evaluation in order to assess model stability.